Recap: gene expression
In 1958, Francis Crick stated the famous “central dogma of molecular biology”, which described the information flow within living cells (Figure 1). He postulated that the information is “stored” chemically in the sequence of DNA, which can be replicated to allow cell division. This information is copied - “transcribed” - onto chemically very similar RNA, which can deliver it from the nucleus to the cytoplasm. The message is then “read” by ribosomes and used to synthesise – or “translate” - the protein, whose nature was “encoded” in it. The whole process of going from the information stored in our DNA to the function exerted by the proteins is what is often referred to as “gene expression”.
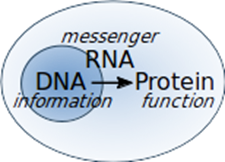
Figure 1. The central dogma of molecular biology
Task
The sequence below is a stretch of mRNA encoding a human protein. “Translate” it computationally into a sequence of amino acids, using EMBOSS Transeq, and find out what this protein is using protein BLAST.
AUGGCGCGUACGAAGCAGACUGCUCGCAAGUCCACCGGCGGCAAGGCUCCGCGCAA
GCAGCUGGCCACCAAGGCGGCUCGGAAGAGCGCUCCGGCCACCGGCGGUGUCAAGA
AGCCCCAUCGCUAUCGGCCUGGUACAGUGGCUCUCCGCGAGAUUCGCCGCUACCAG
AAGUCCACCGAGCUGCUGAUCAGAAAGCUGCCUUUUCAGCGUCUGGUGCGUGAGAU
CGCGCAGGACUUCAAGACCGACUUGCGCUUCCAGAGCUCCGCGGUGAUGGCGCUGC
AAGAGGCAUGCGAGGCCUACCUGGUGGGGCUCUUUGAGGACACCAACCUGUGCGCC
AUCCACGCCAAGCGGGUGACUAUCAUGCCCAAGGACAUCCAGCUCGCACGUCGUAU
CCGCGGCGAGAGGGCUUGA
EMBOSS Transeq online tool: https://www.ebi.ac.uk/Tools/st/emboss_transeq/
This tool takes a sequence of nucleotides and calculates what the sequence of amino acids (i.e. of a protein) encoded by this DNA or RNA would be, using the standard genetic code. Copy the RNA sequence into the query window and click ‘Submit’ to see the result!
NCBI protein BLAST: https://blast.ncbi.nlm.nih.gov/Blast.cgi?PROGRAM=blastp&PAGE_TYPE=BlastSearch&LINK_LOC=blasthome
BLAST is an alignment tool that aligns a given sequence against its vast database of all sequences known to date. Protein BLAST, as the name suggests, does it for various proteins. Copy the amino acid sequence into the input window. Remove the last asterisk that denotes the stop codon (it confuses the program). To narrow the search database down, type and choose ‘human’ in the ‘Organism’ field, then press ‘BLAST’. The output is somewhat complicated. Hide the ‘Graphic Summary’ box by clicking the minus sign next to it, and then check the top entry in the ‘Sequences producing significant alignments’ list. It should have 100% coverage and 100% identity with the query sequence – this is your answer!
Even though the central dogma is still believed to be correct in the first approximation, the picture turned out to be much more complex. In this set of activities you will learn about one of the additional layers complementing the above picture – epigenetics.
What is epigenetics?
To start with, we must understand (or at least try to) what this strange word “epigenetics” means at all. Epigenetics is quite a controversial area of biology, and actually the controversy starts already in the definition, since by saying “epigenetics” people often mean different things, depending on what perspective they look at it from.
Epigenetics and development
The term was first suggested by Conrad Waddington in 1942. Waddington was interested in understanding how a single cell can give rise to a multicellular organism, in which cells are genetically identical and yet display remarkably different features. He defined the “epigenotype” as a complex of developmental processes connecting genotype and phenotype, i.e. how the information in genes is read during embryonic development to give a whole organism. He likened the process of cell differentiation to a marble on top of a hill, that can take one of the downhill paths, leading to a defined terminally differentiated state, and called this model the “epigenetic landscape” (Figure 2).
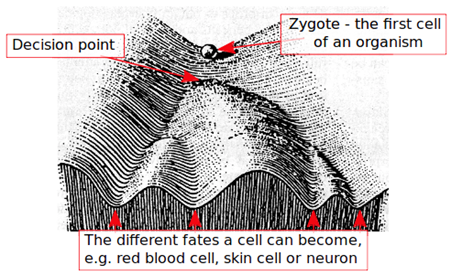
Figure 2. Waddington’s “epigenetic landscape
Epigenetics and gene expression
The variability between differentiated cells is the result of differential gene expression. Hence, to answer Waddington’s question, it is necessary to understand how this expression is regulated and how this “epi-genetic” (i.e. “above genetic” or “beyond genetic”) regulatory information is passed from cell to cell. Therefore, by epigenetics we often mean the inheritance of gene expression patterns across organism or cell generations without changes in the DNA sequence itself.
Epigenetics and molecular biology
Lastly, epigenetics seeks to find out how the information on gene expression patterns can be mechanistically “recorded” and “implemented”. Scientists want to understand what other features on top of the DNA sequence itself can act as information carriers, and decipher the “epigenetic code”. From this mechanistic, molecular point of view, we can define epigenetics as the structural adaptation of chromosomal regions, which results from or leads to altered activity of the genes in this region.
Task
Before going to the next activity, can you guess how the physical structure of a gene and its local environment can influence its expression?


